In an ideal world, scientists would have access to the entire body of published scientific knowledge. They would be able to search this resource to rapidly locate and extract specific data in a way that is accurate, is repeatable, and leads to the discovery of new and related information. We’re not there yet, but technology and information science have made great strides in this direction.
Scientific publications contain measurements, descriptions, and images that have utility beyond the aims of the original work, particularly when they are aggregated into databases. For example, the Paleobiology Database contains field- and museum-based descriptions of more than 1.3 million fossil occurrences compiled from some 50,000 references, and sample-based geochemical data from the published literature are available in EarthChem. Both databases can be used to address fundamental scientific questions, but neither is complete. Plus, adding to existing literature-based data syntheses and constructing new ones is difficult and can be prohibitively time-consuming.
The primary goal of our U.S. National Science Foundation EarthCube building block project, GeoDeepDive, is to facilitate the creation and augmentation of literature-derived databases and to leverage published knowledge and past investments in data acquisition. The project combines library science (the aggregation and curation of digital documents and bibliographic metadata), geoscience (the generation of research questions and labeling of terms in externally managed scientific ontologies), and computer science (the use of high-throughput computing infrastructure and machine reading systems to parse and extract data from millions of documents).
Here we report on the status of this project and describe how the GeoDeepDive infrastructure can be used in scientific research and education applications.
A Next-Generation Digital Library
At least three barriers must be overcome before data can be extracted accurately, repeatably, and with broad applicability from the entire body of published scientific knowledge.
Accurate, repeatable, and broadly applicable extraction of data from the entire body of published scientific knowledge isn’t possible yet. At least three barriers must be overcome before these objectives can be realized.
The first barrier is the time and difficulty involved in assembling millions of published scientific papers into a comprehensive collection that can be used for text and data mining purposes. Open access is a welcome addition to scientific publication, but open access publications remain dispersed across different sources, and providers of open access content have many different protocols for exposing documents and bibliographic metadata. Commercial publishers may produce scientifically relevant publications, but most have infrastructure that restricts document and metadata distribution. GeoDeepDive is addressing these challenges with its own infrastructure that automatically downloads and stores digital documents and bibliographic metadata from many different open source and commercial publishers on an ongoing basis.
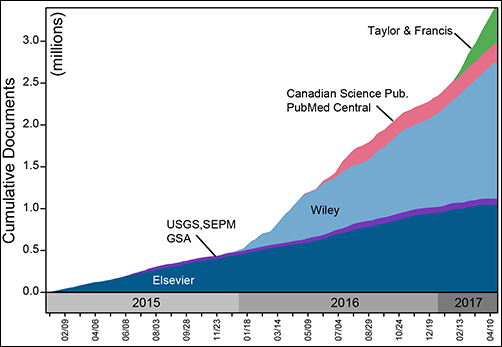
Since our first agreement in 2014, forged between Elsevier and our subscriber institution, the University of Wisconsin–Madison Libraries, GeoDeepDive’s coverage of the published literature has expanded steadily. We cover open access content from the U.S. Geological Survey (USGS), the Public Library of Science (PLoS), and PubMed Central, and we now hold agreements with Wiley, Taylor and Francis, Canadian Science Publishing, the Geological Society of America, and the Society for Sedimentary Geology (Figure 1). Negotiations with other publishers and societies are ongoing. Geoscience remains our disciplinary focus, but the document acquisition and processing infrastructure is agnostic with respect to discipline: All scientific and biomedical fields are covered.
The second barrier to leveraging the published literature is the computing capacity required to parse and analyze millions of documents. Storing files locally allows us to leverage HTCondor software and high-throughput computing to parse and annotate documents rapidly. We use a variety of software tools to process original content. Stanford CoreNLP, a natural language processing toolkit, parses text into sentences, parts of speech, and linguistic dependencies (Figure 2). Tesseract-ocr, an optical character recognition (OCR) tool, parses documents by layout elements so that we can identify and extract figures and tables. Elasticsearch is used to locate terms and phrases of interest derived from externally constructed and curated hierarchical dictionaries.
Users of GeoDeepDive do not have direct access to original documents. Instead, they have access to output generated by millions of computing hours devoted to parsing millions of published papers. Full citation data, along with URLs pointing back to original sources, are always available.

The third barrier to leveraging the published literature is the time and effort that is required to develop software tools that can recognize and extract data and information that are relevant to a given problem. What’s more, this data extraction must meet the data quality standards demanded by scientists. There is currently no single solution to this problem. Instead, the goal of GeoDeepDive is to make the development of useful applications as easy and efficient as possible.
Developing Custom Applications and Using GeoDeepDive
To develop a GeoDeepDive application, users must write code that reads and analyzes the output of natural language processing (NLP), OCR, or custom document parsing and annotation tools.
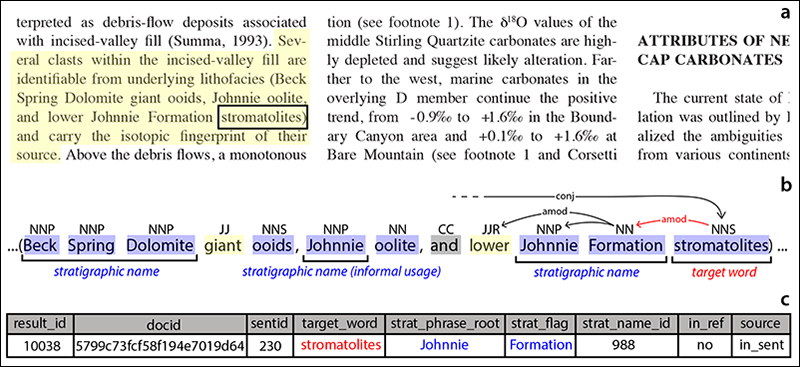
Undergraduate student Julia Wilcots wrote the first application to use GeoDeepDive infrastructure [Wilcots et al., 2015]. The original application, modified by Peters et al. [2017] and available on GitHub in the Python computer language, first identifies all sentences containing the word “stromatolite” (laminated, microbially formed sedimentary structures) and its variants (Figure 3). It then locates stratigraphic names (e.g., “Belt Supergroup”) within that same sentence or in adjacent sentences. The output is a list of individual mentions of stromatolites, the rock units that contain them, and a citation or link to the relevant publication.
Another research intern, Erika Ito, is developing an application to investigate the completeness of the Paleobiology Database relative to the published literature. This app, written in the R language, uses NLP output to locate mentions of geologic formations that are described as fossil bearing but that are not currently in the Paleobiology Database. The end result is a machine-executed assessment of how thoroughly paleontologists have collated the scientific literature. The application also produces a priority list for future data acquisition efforts.
GeoDeepDive allows one-click discovery of publications that are potentially relevant to users who are exploring geological maps in the field.
GeoDeepDive infrastructure can also be used to provide links to published documents that contain specific terms and phrases of interest. For example, the Macrostrat database uses the Web-based GeoDeepDive application programming interface (API) to link lithostratigraphic names in geologic maps to the published literature (Figure 4). When users click on named rock units in a Web browser or tap on named map units in a mobile app, the GeoDeepDive API retrieves snippets of text around mentions of those same terms in the full text of documents, along with citation metadata and document URLs. This allows one-click discovery of publications that are potentially relevant to users who are exploring geological maps in the field.

Individuals wishing to explore how GeoDeepDive can be used in their projects have several options. The API is open and offers some useful capabilities right now, such as the ability to explore journal coverage and retrieve links to documents that contain specific terms. New dictionaries can be ingested at any time, and we are eager to involve individuals and communities who have structured vocabularies that can be indexed against the literature. Several basic documentation-type tutorials and example apps are also available on GitHub (Figure 5).

Next Steps
Development efforts to date have focused on forging agreements with publishers and improving GeoDeepDive infrastructure for downloading, storing, parsing, and annotating documents. User applications have focused on text-based information [e.g., Liu et al., 2016; Peters et al., 2017], but a large amount of data resides in tables and figures. We are working to allow users to write applications that read the full text of documents, identify specific tables and figures of interest, and then extract data from them.
We are still far from the pie-in-the-sky capabilities that are often imagined in the context of machine reading and learning applied to the scientific literature. However, GeoDeepDive is establishing a solid building block for the future by focusing on building a comprehensive digital library, backed by agreements and partnerships with publishers and content owners, that is supported by a reliable high-throughput computing infrastructure. We look forward to helping geoscientists to more efficiently discover and leverage hard-earned data locked in the scientific literature.
Acknowledgment
This work was supported by NSF EarthCube ICER 1343760.
References
Corsetti, F. A., and A. J. Kaufman (2003), Stratigraphic investigations of carbon isotope anomalies and Neoproterozoic ice ages in Death Valley, California, Geol. Soc. Am. Bull., 115, 916–932,https://doi.org/10.1130/B25066.1.
Liu, C., et al. (2016), Reconstructing the evolution of first-row transition metal minerals by GeoDeepDive, Abstract IN41A-1649 presented at 2016 Fall Meeting, AGU, San Francisco, Calif.
Peters, S. E., J. M. Husson, and J. Wilcots (2017), Rise and fall of stromatolites in shallow marine environments, Geology, 45, 487, https://doi.org/10.1130/G38931.1.
Wilcots, J., J. Husson, and S. E. Peters (2015), Stromatolite distribution in space and time: A machine-reading assisted quantitative analysis, Geol. Soc. Am. Abstr. Programs, 47, 365.
Author Information
Shanan E. Peters (email: [email protected]), Department of Geoscience, University of Wisconsin–Madison; Ian Ross, Department of Computer Sciences, University of Wisconsin–Madison; John Czaplewski, Department of Geoscience, University of Wisconsin–Madison; Aimee Glassel, University Libraries, University of Wisconsin–Madison; Jon Husson, Valerie Syverson, and Andrew Zaffos, Department of Geoscience, University of Wisconsin–Madison; and Miron Livny, Department of Computer Sciences, University of Wisconsin–Madison
Citation:
Peters, S. E.,Ross, I.,Czaplewski, J.,Glassel, A.,Husson, J.,Syverson, V.,Zaffos, A., and Livny, M. (2017), A new tool for deep-down data mining, Eos, 98, https://doi.org/10.1029/2017EO082377. Published on 22 September 2017.
Text © 2017. The authors. CC BY-NC-ND 3.0
Except where otherwise noted, images are subject to copyright. Any reuse without express permission from the copyright owner is prohibited.

