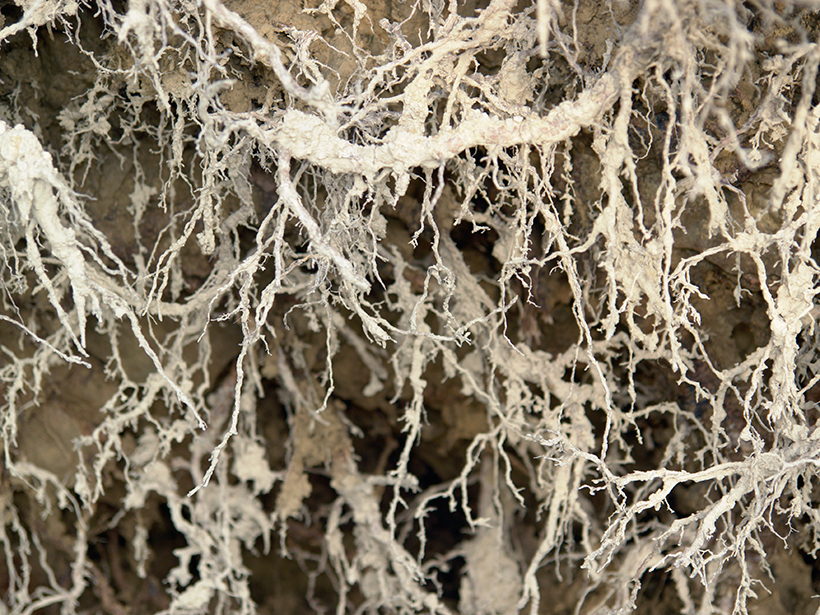
Fine roots are complex and dynamic structures that serve many critical roles for land plants. Their well-known importance for acquiring water and nutrients and associations with soil microbes underpin many terrestrial processes, including the movement of carbon, nutrients, and water through terrestrial ecosystems.

Despite this importance, uncertainty shrouds the traits and processes associated with these enigmatic organs. Such uncertainty limits confidence in our terrestrial biosphere models that incorporate information based on fine-root data. For example, without knowledge of how roots grow and function, scientists struggle to predict how plants and ecosystems will respond to changes in atmospheric carbon dioxide as belowground resources become more limiting to plant growth.
To address this challenge, an international group of researchers developed and released in 2017 the first version of a new resource: the Fine-Root Ecology Database (FRED) [Iversen et al., 2017]. Since that time, FRED has undergone significant improvement and additions, culminating in the release of FRED version 2.0 [Iversen et al., 2018]. FRED is a fully open resource containing species-specific as well as mixed-species root trait data that encompass all vegetated terrestrial biomes.
Meet FRED
Data collection and curation are focused on fine roots, but FRED also includes data on coarse roots, total belowground biomass, and associations between roots and microbes.
The representation of root traits in most terrestrial models leaves substantial room for improvement. For example, the rate at which fine roots grow, die, and get replaced can vary by more than an order of magnitude between models that represent generic temperate forest plant functional types [McCormack et al., 2015]. At the same time, the Community Land Model assigns a fixed value to the fine-root carbon-to-nitrogen ratio across all global biomes—from savanna grasses to tropical rain forests—without any variation for plant functional types [Sargsyan et al., 2014].
The coarseness of model representations for many root-related processes stems largely from a lack of robust data to constrain parameter estimates and derive empirical relationships among root traits and between root and ecosystem functions.
FRED 2.0 provides a resource to address these issues. The database is built on data collected from primary literature, in addition to direct data contributions, and it is freely available for download without restriction. This latest version release contains more than 100,000 trait observations from roughly 300 traits (such as root diameter and root respiration), among eight distinct trait types (such as morphology and physiology; described in McCormack et al. [2017]).

Data collection and curation are focused on fine roots, but FRED also includes data on coarse roots, total belowground biomass, and associations between roots and microbes. Trait observations are recorded along with relevant ancillary data that include site location, climate, and soil characteristics. We also used external data products to assign plant functional type and biome classifications to each observation. Finally, we used a common taxonomy to catalog species-specific data observations in FRED 2.0.
In recent years, scientists have increasingly recognized that the traditionally defined fine-root pool (i.e., all roots ≤ 2 millimeters in diameter) includes a heterogeneous mixture of roots with diverse form and function. Making all trait measurements on a mixed fine-root pool is akin to analyzing a sample of leaves and woody branches all jumbled together. Accordingly, FRED is explicitly structured to organize trait observations made on different classes of fine roots, enabling meaningful trait comparisons and better parameterization of roots in models.
Working with FRED
Empiricists and modelers interested in downloading FRED 2.0 can access the full database, which provides three files for download.
First, a user guidance document provides information on data acquisition and guidelines for data use, as well as a data dictionary, which is a table of the traits included within the database together with all relevant abbreviations and trait definitions. In particular, each trait has been assigned a unique identifier. The identifier, referred to as Column ID in the guidance document, is searchable within the database to avoid confusion among trait listings that may differ only in their method of measurement or units. This document also specifies our data use and privacy policies regarding user data and presents a series of tips for how to effectively use FRED.
Second, we provide a full list of all data sources used to compile FRED. Finally, the entire database is available for download as a single CSV file.
In addition to these files, the FRED website is a good source of information. There, we further provide searchable lists of root traits and ancillary data. We also provide a list of all plant species included in the database to help users determine whether particular sets of information are currently housed in FRED before they download the full database.
We also provide tips on the best use of the observations housed within FRED for users who may not be familiar with the interpretation and use of fine-root data. Users interested in fine-root observations can also refer to the extensive bibliography of data sources included in FRED, and this bibliography further serves as a check to see whether particular project data have been incorporated into FRED.
FRED’s Future
We are continuing our efforts to expand and improve the database coverage and accessibility. As part of these efforts, we encourage feedback from modelers working to update parameterizations and descriptions of root-related processes in various modeling applications. Such feedback might include conceptual representations of how roots and root functions would be idealized in their model structures. These qualitative assertions could then become a valuable tool to inform future trait measurements in the field and direct our efforts incorporating data into the next version of FRED.
Better connections between empirical research and physiological processes are one area that is ripe for improvement and that will be particularly important for future model developments. Easily measured root traits, including morphology, chemistry, and anatomy, must be connected to the physiological processes that underpin root function in both the natural world and the models that represent this world.
Reports of common empirical traits like root nitrogen content, root diameter, and standing root biomass in FRED far outpace available measurements of dynamic processes like root respiration, resource uptake, and life span. Yet it is precisely these latter metrics that are needed to determine movement of carbon, nutrients, and water in terrestrial ecosystems.
In the near term, researchers will be able to use the data now available in FRED 2.0 to derive statistical and, where possible, mechanistic relationships among these traits to better estimate process rates for roots in models. Over the longer term, however, more targeted campaigns that explicitly measure the root functions and belowground processes represented in terrestrial biosphere models will help to solidify our empirical knowledge base and validate model parameterizations.
Acknowledgments
C.M.I., M.L.M., and A.S.P. were supported by the Office of Biological and Environmental Research in the U.S. Department of Energy (DOE) Office of Science. This manuscript has been authored by UT-Battelle, LLC under contract DE-AC05-00OR22725 with the DOE. The U.S. government retains and the publisher, by accepting the article for publication, acknowledges that the United States government retains a nonexclusive, paid-up, irrevocable, worldwide license to publish or reproduce the published form of this manuscript, or allow others to do so, for U.S. government purposes. DOE will provide public access to these results of federally sponsored research in accordance with the DOE Public Access Plan.
References
Iversen, C. M., et al. (2017), A global Fine-Root Ecology Database to address below-ground challenges in plant ecology, New Phytol., 215(1), 15–26, https://doi.org/10.1111/nph.14486.
Iversen, C. M., et al. (2018), Fine-Root Ecology Database (FRED): A global collection of root trait data with coincident site, vegetation, edaphic, and climatic data, version 2, Oak Ridge Natl. Lab. Terrestrial Ecosyst. Sci. Sci. Focus Area, Oak Ridge, Tenn.
McCormack, M. L., et al. (2015), Sensitivity of four ecological models to adjustments in fine root turnover in temperate forests: Does fine root turnover matter?, Ecol. Modell., 297, 107–117, https://doi.org/10.1016/j.ecolmodel.2014.11.013.
McCormack, M. L., et al. (2017), Building a better foundation: Improving root-trait measurements to understand and model plant and ecosystem processes, New Phytol., 215(1), 27–37, https://doi.org/10.1111/nph.14459.
Sargsyan, K., et al. (2014), Dimensionality reduction for complex models via Bayesian compressive sensing, Int. J. Uncertainty Quantif., 4(1), 63–93, https://doi.org/10.1615/Int.J.UncertaintyQuantification.2013006821.
Author Information
M. Luke McCormack (email: [email protected]), Department of Plant and Microbial Biology, University of Minnesota Twin Cities, St. Paul; and A. Shafer Powell and Colleen M. Iversen, Climate Change Science Institute and Environmental Sciences Division, Oak Ridge National Laboratory, Tenn.
Citation:
McCormack, M. L.,Powell, A. S., and Iversen, C. M. (2018), Better plant data at the root of ecosystem models, Eos, 99, https://doi.org/10.1029/2018EO104093. Published on 21 August 2018.
Text © 2018. The authors. CC BY 3.0
Except where otherwise noted, images are subject to copyright. Any reuse without express permission from the copyright owner is prohibited.