Researchers on the cutting edge of geochemistry are tracing Earth’s history through the clues recorded in tiny mineral crystals. Often, they must work with the microscopic (or nanoscopic) features in rare, hard-won specimens to pull apart the complete history of the mineral’s formation. Even in large pieces of rock, microscopic variability may be distributed throughout in patterns that contain valuable information. Researchers using microanalysis have read the information stored in layers of a single zircon crystal, the cements in a sandstone, tiny shells of sea creatures like ammonite hatchlings, and minerals in a meteorite.
A variety of imaging and chemical analysis techniques produce maps illustrating how chemical elements and crystal growth features are distributed across a specimen. Currently, these data are shared through static figures or images contained in a data repository, but this setup does not facilitate deeper inquiry and full use of spatial relationships. What if widely used geographic information system (GIS) software, which typically deals with data on the scale of many square kilometers, could be applied to data on a microscopic scale?
Here we showcase use of the free and open source software QGIS to integrate in situ chemical data with images to enrich interpretation.
Integrating the Information
Minimally destructive geochemical analysis at the nanometer to micrometer scale can peel apart the events that took place over the formation of a mineral crystal and still preserve the crystal for future analyses.
Minimally destructive geochemical analysis at the nanometer to micrometer scale can peel apart the events that took place over the formation of a mineral crystal and still preserve the crystal for future analyses. This analysis is especially important for deciphering the geological history of a specimen that is small, rare, or zoned, like a 4.4-billion-year-old zircon.
No one analytical technique, however, can provide all necessary information for many questions, and the range of instrument manufacturers and data types involved makes integrating data from various instruments difficult.
To understand, for instance, how cementation in a sandstone evolved or the chemical evolution of a magma, images from light microscopes or scanning electron microscopes (SEM) are used to examine petrographic relationships, and then electron probe microanalysis (EPMA) provides point analyses of elemental composition (Figure 1).
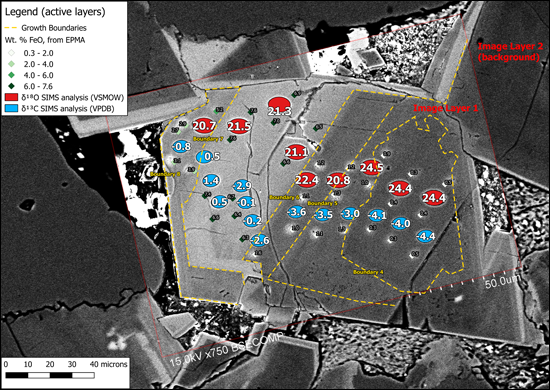
In situ microanalytical techniques have become faster and more precise, and they are now routinely used in the geosciences [Valley and Kita, 2009; Sylvester and Jackson, 2016]. Microanalytical instruments like secondary ion mass spectrometers (SIMS), electron probe microanalyzers, and laser ablation–inductively coupled plasma–mass spectrometers produce large data sets from individual thin sections or chips (Figure 1). Other techniques like optical light microscopy, SEM, X-ray mapping, and nano-SIMS produce suites of images.
A Change of Scale
For years, scientists have managed field-scale data that span scales of meters to kilometers, incorporating images and point information using GIS software. However, in situ microanalytical data, like SEM images and SIMS or EPMA point analysis, also fundamentally contain spatial information that can be combined using GIS.
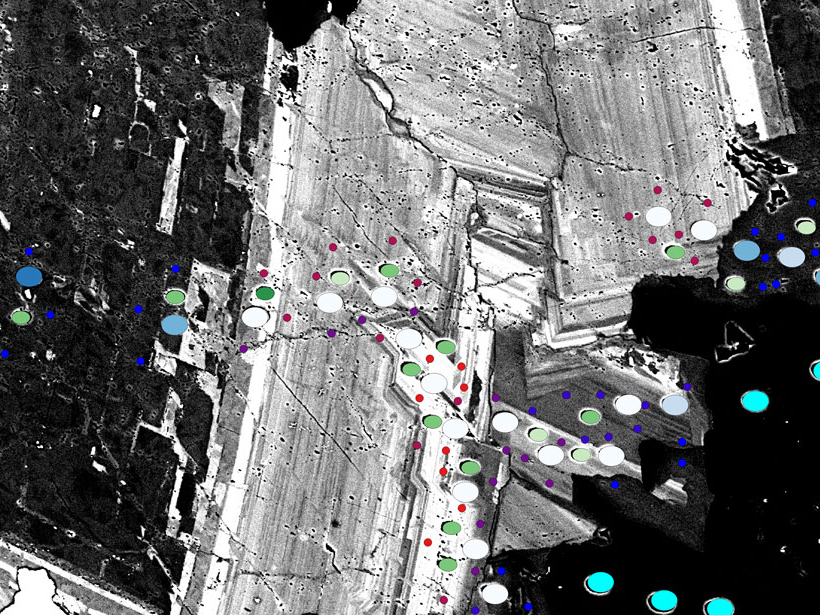
On a field scale, data in a map may have vector components (e.g., points, lines, or polygons) or raster components (e.g., images or similar pixel-based scans). In the case of microanalytical data, SIMS or EPMA data can be treated as vectors, and SEM scans or other images can be treated as rasters. Spatially overlapping sampling images and point analyses allow relation of these data sets to each other for visualization and further exploration.
Adapting an Open Source Tool
QGIS is a free, open source, user-friendly, cross-platform (Mac, Windows, Linux, Unix, and Android) software package for collecting and processing macroscopic geographic data. Under the banner of the Open Source Geospatial Foundation, a community of volunteers maintains and develops the software with financial support from corporate and nonprofit sponsors.
We have compiled in situ microanalytical data from multiple instruments to create integrated maps of individual samples.
Although QGIS was not originally designed for microanalytical techniques, we have compiled in situ microanalytical data from multiple instruments to create integrated maps of individual samples.
QGIS has many advantages, making it a good candidate for adoption by the wider microanalytical community for data visualization and management. First, the open-source nature removes the cost barrier to adopting the platform. Second, the software is supported by a growing international community of users and developers. Finally, custom plug-ins can be written in the Python, C++, and R computer languages, and these can provide ways of refining data input for different instruments. Plug-ins written in Python can be tailored to specific structured instrumental outputs to speed data compilation.
Map generation in QGIS for microanalytical data sets is straightforward (Figure 2). A tutorial structured to teach refined use of QGIS is available at the WiscSIMS Micro-QGIS website.
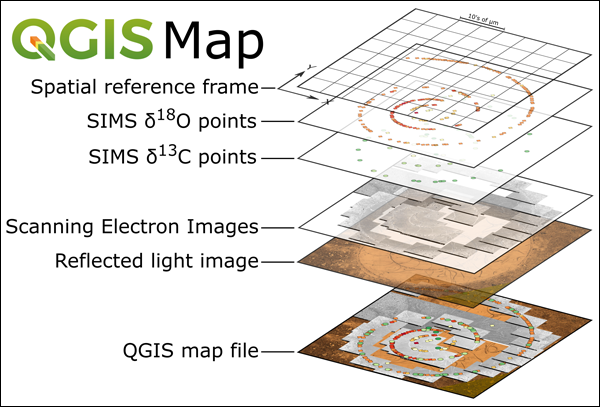
Creating Plug-ins
QGIS plug-ins can be created to solve specific problems relevant to a lab, analytical technique, or scientific question. We have used a plug-in called Plugin Builder to create several tools specific to the WiscSIMS laboratory at the University of Wisconsin–Madison. This platform provides a basic framework and files for building your own plug-in to interface with QGIS.
We have developed a plug-in that places images using the x–y coordinates of a microscope stage to align SEM images with data maps from other instruments. The software parses a text file made by our SEM (Hitachi S3400-N) that contains image dimension, stage coordinates, and pixel size and loads the associated images into a QGIS map. We are distributing our SEM Image Placer plug-in on GitHub, so anyone can download, fork, modify, and make a pull request for the development under the GNU General Public License to extend this functionality to other types of SEMs and other microscopes with stage coordinates.
Integrating and Archiving a Diversity of Data
Using free, open source QGIS software for the integration of in situ microanalytical data sets improves our ability to integrate data from multiple instruments. It also decreases the time spent annotating and reformatting supplemental data sets for publication (Figures 1 and 3).
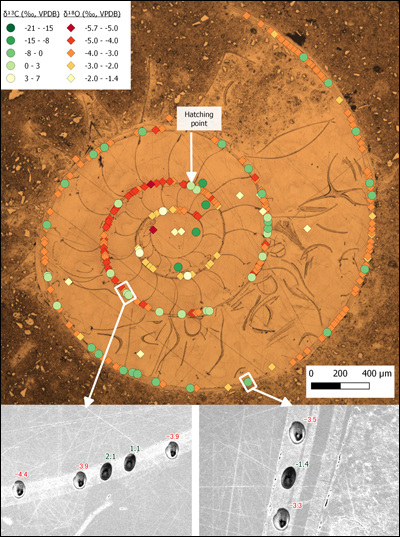

Data collected from multiple instruments can be compiled quickly using built-in tools and custom-designed components. Adapting GIS software to the microscale also enables us to perform more complex spatial analysis. Most important, these compiled microspatial data can be shared between collaborators and used to explore patterns that may not be obvious from static displays of images and data points.
Our compilations use standard, widely used file formats, so they have the potential to interface easily with database infrastructure for archiving and depositing data collected with public funding [Chan et al., 2016]. This potential will ensure that extensive, expensive, and cutting-edge in situ geochemical data sets are easily shared and integrated into existing databases like Macrostrat using widely available formats and software.
Acknowledgments
We thank John Czaplewski, Shanan Peters, Ian Orland, and others for discussion and troubleshooting the QGIS workflow. The WiscSIMS laboratory is partially supported by the National Science Foundation (EAR03-19230, EAR13-55590) and the University of Wisconsin–Madison.
References
Cammack, J. N. (2015), SIMS microanalysis of the Strelley Pool Formation cherts and the implications for the secular-temporal oxygen-isotope trend of cherts, M.S. thesis, University of Wisconsin–Madison, Madison.
Cammack, J. N., et al. (2018), SIMS microanalysis of the Strelley Pool Formation cherts and the implications for the secular-temporal oxygen-isotope trend of cherts, Precambrian Res., 304, 125–139, https://doi.org/10.1016/j.precamres.2017.11.005.
Chan, M. A., S. E. Peters, and B. Tikoff (2016), The future of field geology, open data sharing and cybertechnology in Earth science, Sediment. Rec., 14, 4–10, https://doi.org/10.2110/sedred.2016.1.
Denny, A. C., et al. (2017), Isotopically zoned carbonate cements in early Paleozoic sandstones of the Illinois Basin: δ18O and δ13C records of burial and fluid flow, Sediment. Geol., 361, 93–110, https://doi.org/10.1016/j.sedgeo.2017.09.004.
Linzmeier, B. J., et al. (2018), Ion microprobe measured stable isotope evidence for the habitat and mode of life of ammonites in early ontogeny, Paleobiology, in press.
Sylvester, P. J., and S. E. Jackson (2016), A brief history of laser ablation inductively coupled plasma mass spectrometry (LA–ICP–MS), Elements, 12, 307–310, https://doi.org/10.2113/gselements.12.5.307.
Valley, J. W., and N. T. Kita (2009), In situ oxygen isotope geochemistry by ion microprobe, in Secondary Ion Mass Spectrometry in the Earth Sciences, MAC Short Course, vol. 41, edited by M. Fayek, pp. 19–63, Mineral. Assoc. of Can., Quebec.
Author Information
B. J. Linzmeier (email: [email protected]), Department of Earth and Planetary Science, Northwestern University, Evanston, Ill.; K. Kitajima, WiscSIMS Facility, University of Wisconsin–Madison; A. C. Denny, Department of Geoscience, University of Wisconsin–Madison; and J. N. Cammack, Department of Geoscience, University of Wisconsin–Madison; also at Geology Department, Fort Lewis College, Durango, Colo.
Citation:
Linzmeier, B. J.,Kitajima, K.,Denny, A. C., and Cammack, J. N. (2018), Making maps on a micrometer scale, Eos, 99, https://doi.org/10.1029/2018EO099269. Published on 17 May 2018.
Text © 2018. The authors. CC BY 3.0
Except where otherwise noted, images are subject to copyright. Any reuse without express permission from the copyright owner is prohibited.

